ggplot(data = rd) +
geom_line(mapping = aes(x = MESS_DATUM, y = RS, color = STATION))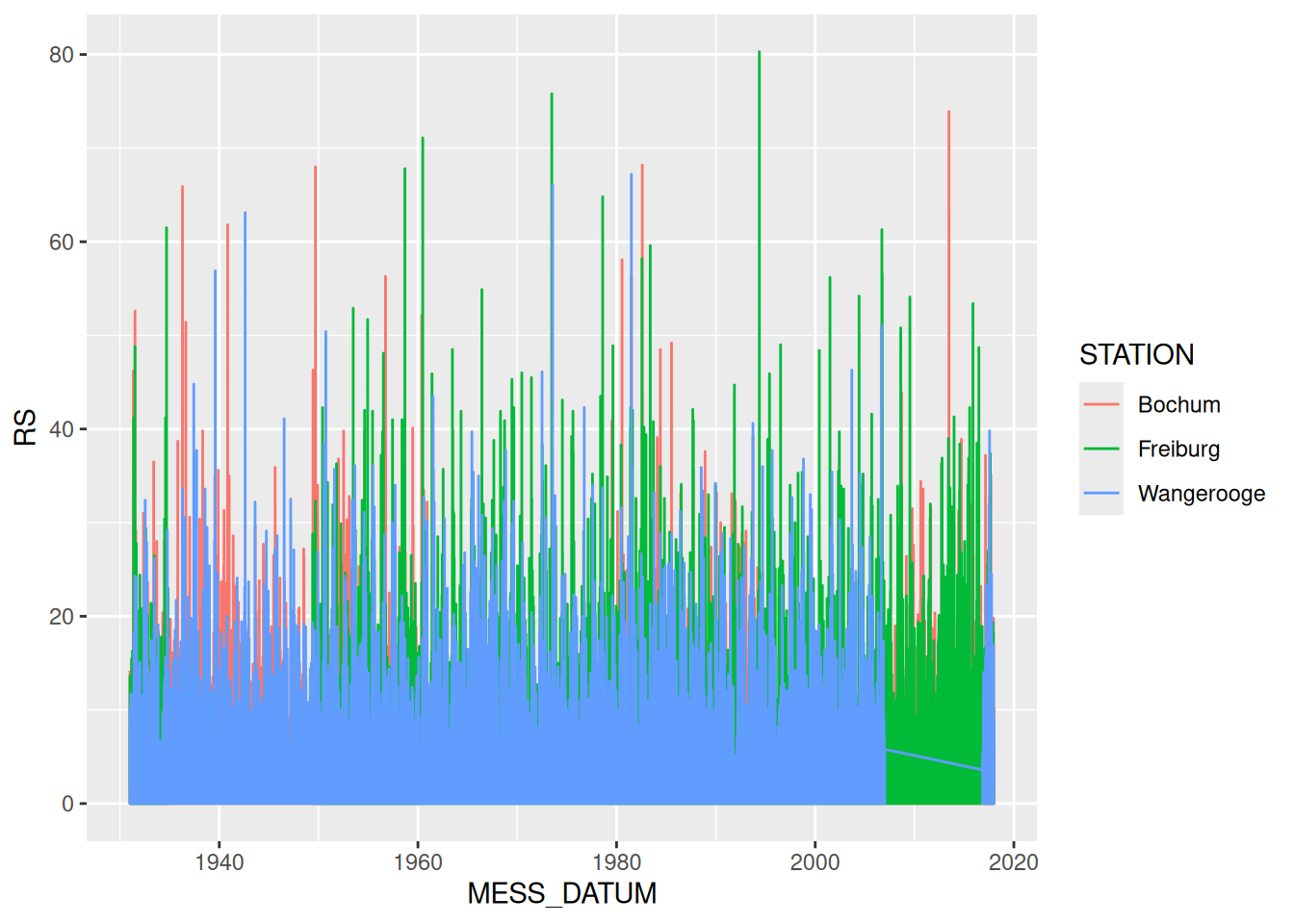
ggplot(data = rd) +
geom_line(mapping = aes(x = MESS_DATUM, y = RS, color = STATION))
scripts/prepare-data.R
rd_freiburg <- rd |> filter(STATION == "Freiburg")
mod_freiburg <- lm(data = rd_freiburg, RS ~ MESS_DATUM)
rd_freiburg <- rd_freiburg |> add_predictions(mod_freiburg) |> as_tibble()
rd_freiburg# A tibble: 26,537 × 4
STATION MESS_DATUM RS pred
<chr> <date> <dbl> <dbl>
1 Freiburg 1931-01-01 5.3 2.56
2 Freiburg 1931-01-02 5.6 2.56
3 Freiburg 1931-01-03 2.4 2.56
4 Freiburg 1931-01-04 13.6 2.56
5 Freiburg 1931-01-05 0 2.56
6 Freiburg 1931-01-06 0 2.56
7 Freiburg 1931-01-07 0.2 2.56
8 Freiburg 1931-01-08 0 2.56
9 Freiburg 1931-01-09 0 2.56
10 Freiburg 1931-01-10 0 2.56
# ℹ 26,527 more rows
lm(data = d, Y ~ X)add_predictions() werden die Werte des Modells hinzugefügtpred (prediction = Vorhersage)add_predictions() aus Paket modelrggplot(data = rd_freiburg) +
geom_line(mapping = aes(x = MESS_DATUM, y = pred), color = "red")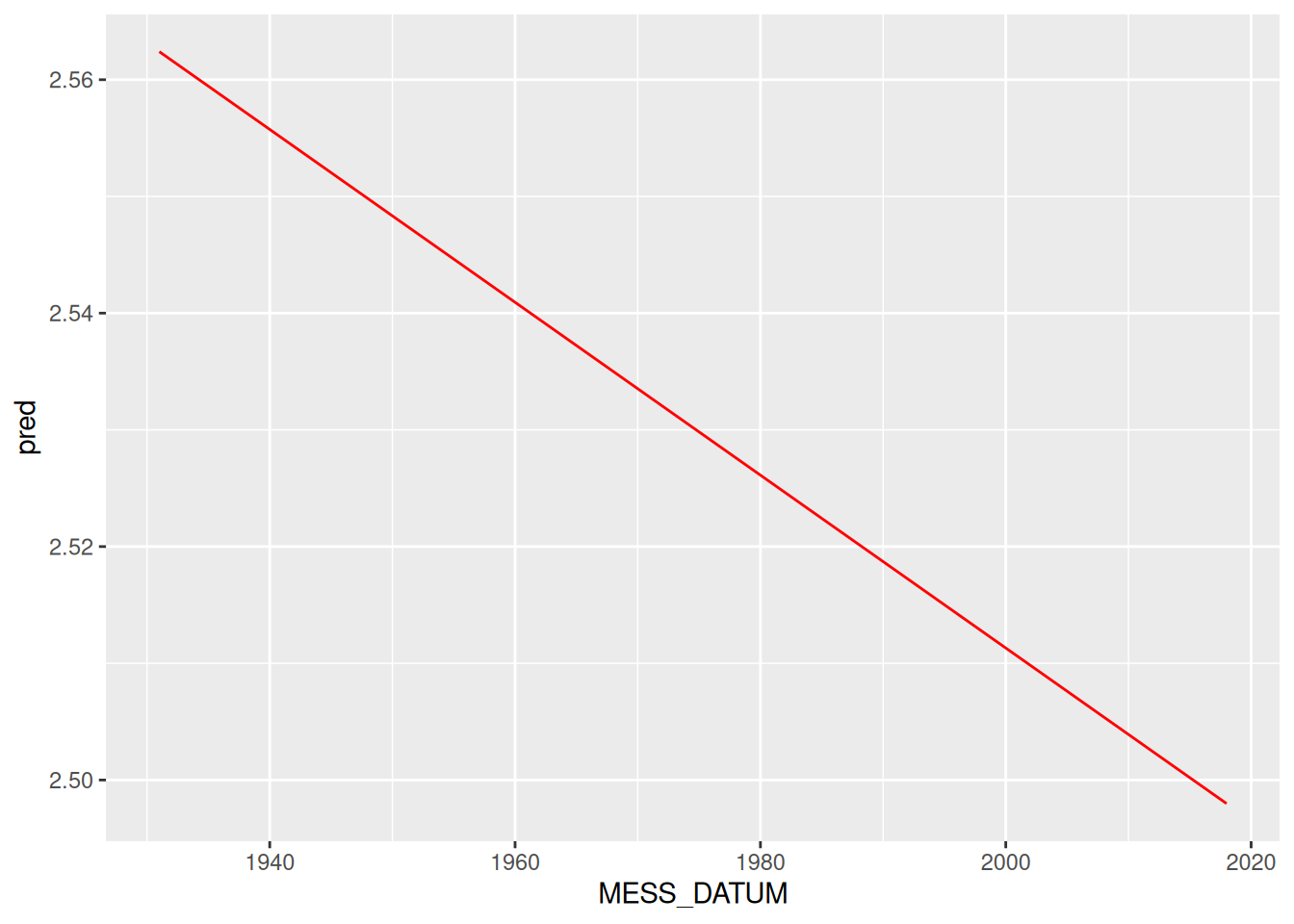
geom_linegeom_smoothrd_stationen <- rd |>
group_by(STATION) |>
nest()
rd_stationen# A tibble: 3 × 2
# Groups: STATION [3]
STATION data
<chr> <list>
1 Wangerooge <tibble [28,093 × 2]>
2 Freiburg <tibble [26,537 × 2]>
3 Bochum <tibble [20,200 × 2]>
data sind jetzt alle Beobachtungen für eine Station zusammengefasstrd_stationen$data[[1]]# A tibble: 28,093 × 2
MESS_DATUM RS
<date> <dbl>
1 1931-01-01 2.9
2 1931-01-02 0.2
3 1931-01-03 7.8
4 1931-01-04 10.3
5 1931-01-05 1.3
6 1931-01-06 0
7 1931-01-07 0
8 1931-01-08 0
9 1931-01-09 0.6
10 1931-01-10 0
# ℹ 28,083 more rows
mapstations_modell <- function(df) { lm(data = df, RS ~ MESS_DATUM) }
map(rd_stationen$data, stations_modell)[[1]]
Call:
lm(formula = RS ~ MESS_DATUM, data = df)
Coefficients:
(Intercept) MESS_DATUM
2.141e+00 1.094e-05
[[2]]
Call:
lm(formula = RS ~ MESS_DATUM, data = df)
Coefficients:
(Intercept) MESS_DATUM
2.534e+00 -2.027e-06
[[3]]
Call:
lm(formula = RS ~ MESS_DATUM, data = df)
Coefficients:
(Intercept) MESS_DATUM
2.215e+00 6.409e-06 stations_modell erstellt ein lineares Modell für einen Dataframemap aus dem Paket purrr erstellen wir ein Modell für jede Stationrd_stationen <- rd_stationen |>
mutate(
MODELL = map(data, stations_modell)
)
rd_stationen# A tibble: 3 × 3
# Groups: STATION [3]
STATION data MODELL
<chr> <list> <list>
1 Wangerooge <tibble [28,093 × 2]> <lm>
2 Freiburg <tibble [26,537 × 2]> <lm>
3 Bochum <tibble [20,200 × 2]> <lm>
MODELL enthält nun die linearen Modellerd_stationen <- rd_stationen |>
mutate(data2 = map2(data, MODELL, add_predictions))
rd_stationen# A tibble: 3 × 4
# Groups: STATION [3]
STATION data MODELL data2
<chr> <list> <list> <list>
1 Wangerooge <tibble [28,093 × 2]> <lm> <tibble [28,093 × 3]>
2 Freiburg <tibble [26,537 × 2]> <lm> <tibble [26,537 × 3]>
3 Bochum <tibble [20,200 × 2]> <lm> <tibble [20,200 × 3]>
data2 enthält jetzt die Daten zu den linearen Trendsmap2 übergibt die beiden Parameter data und MODELL an die Funktion add_predictionsrd_lineare_trends <- rd_stationen |>
unnest(data2) |>
select(-data, -MODELL)
rd_lineare_trends# A tibble: 74,830 × 4
# Groups: STATION [3]
STATION MESS_DATUM RS pred
<chr> <date> <dbl> <dbl>
1 Wangerooge 1931-01-01 2.9 1.99
2 Wangerooge 1931-01-02 0.2 1.99
3 Wangerooge 1931-01-03 7.8 1.99
4 Wangerooge 1931-01-04 10.3 1.99
5 Wangerooge 1931-01-05 1.3 1.99
6 Wangerooge 1931-01-06 0 1.99
7 Wangerooge 1931-01-07 0 1.99
8 Wangerooge 1931-01-08 0 1.99
9 Wangerooge 1931-01-09 0.6 1.99
10 Wangerooge 1931-01-10 0 1.99
# ℹ 74,820 more rowsunnest werden die Dataframes wieder in Zeilen umgewandeltggplot(data = rd_lineare_trends) +
geom_line(mapping = aes(x = MESS_DATUM, y = pred, color = STATION))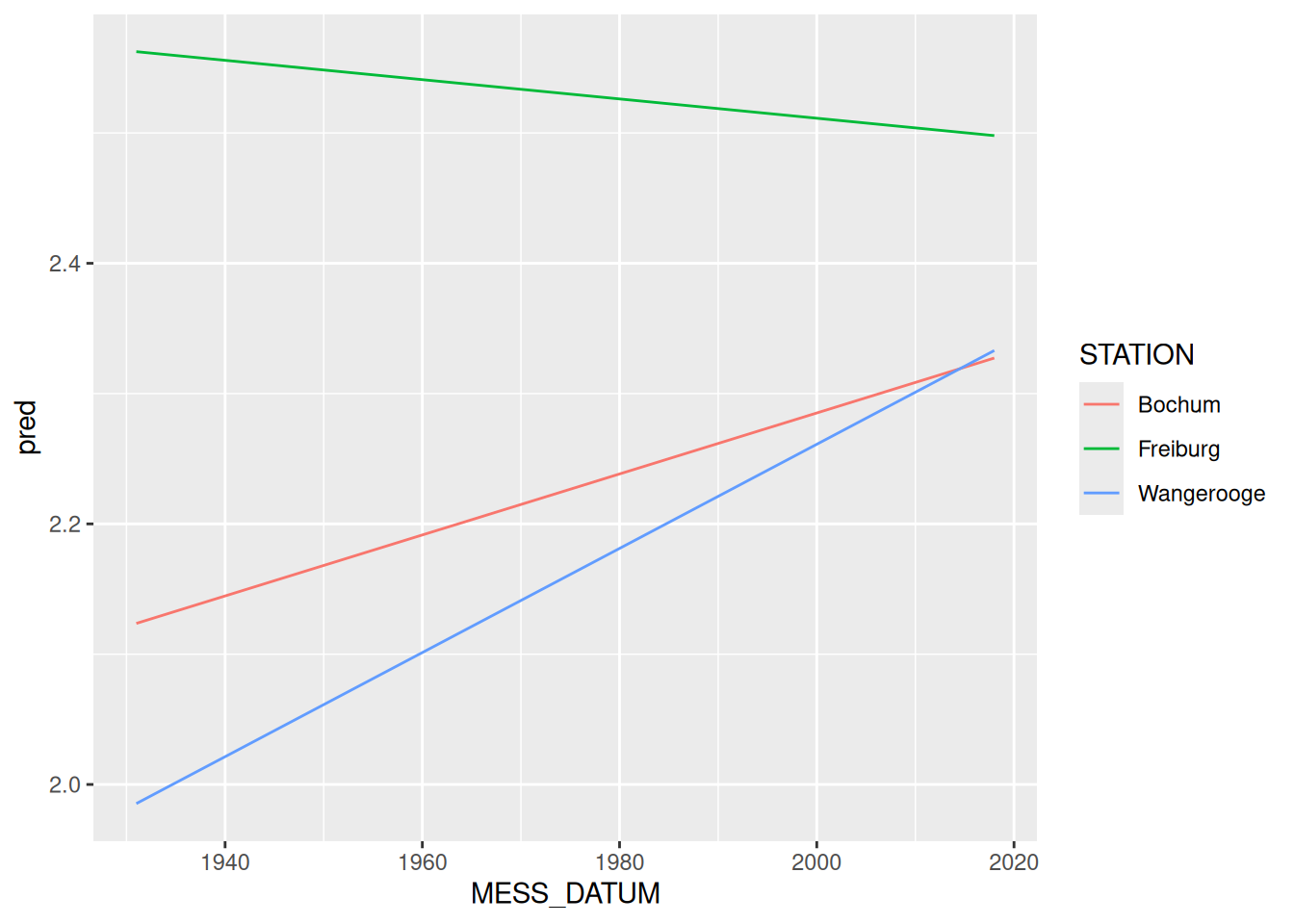
linearer_trend_dataframe <- function(s, df) {
m <- lm(data = df, RS ~ MESS_DATUM)
tibble(STATION = s, ALPHA = m$coefficients[1], BETA = m$coefficients[2])
}
rd_lineare_trends_2 <- map2_dfr(rd_stationen$STATION, rd_stationen$data, linearer_trend_dataframe)
rd_lineare_trends_2# A tibble: 3 × 3
STATION ALPHA BETA
<chr> <dbl> <dbl>
1 Wangerooge 2.14 0.0000109
2 Freiburg 2.53 -0.00000203
3 Bochum 2.21 0.00000641
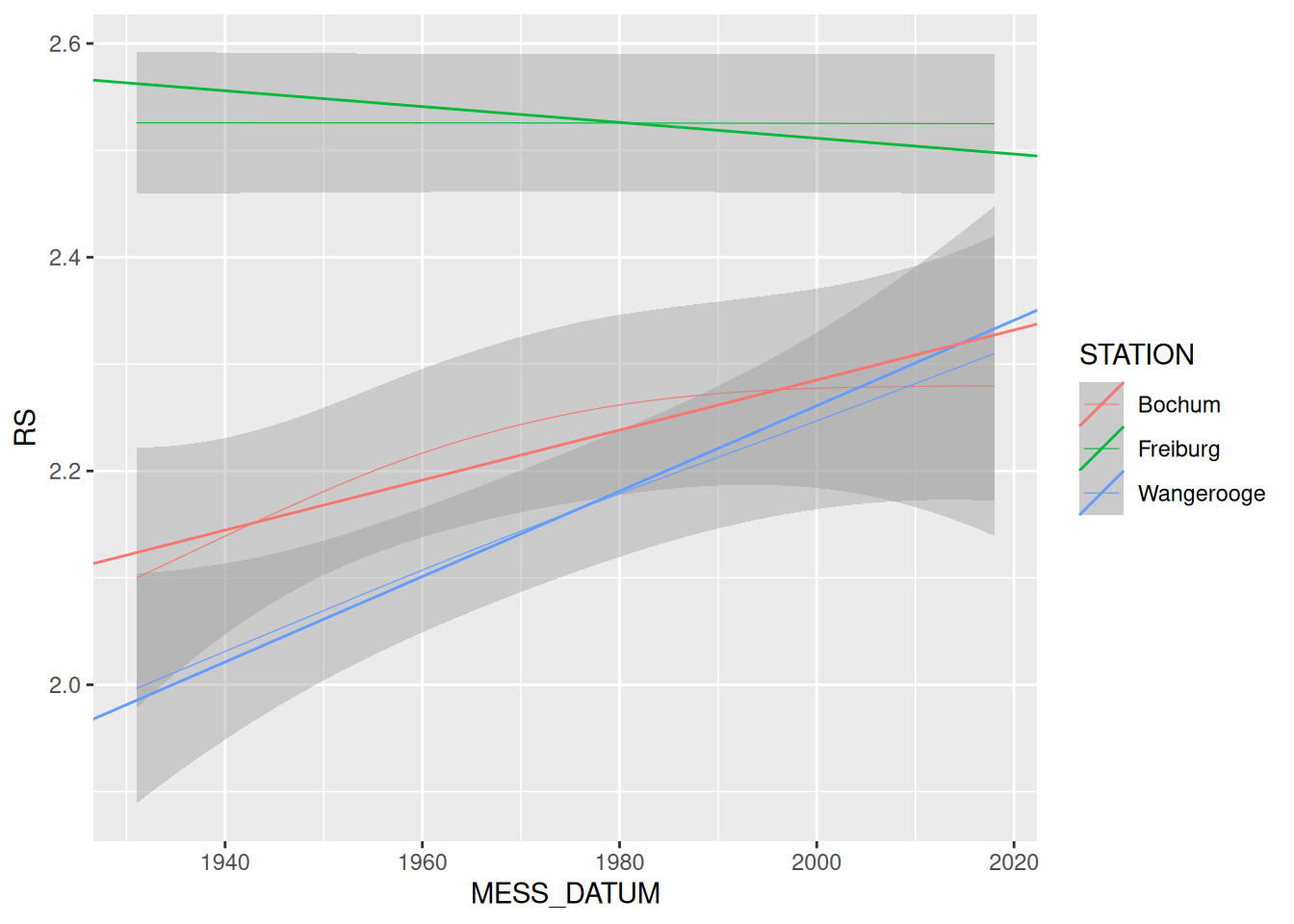
linearer_trend_dataframe erstellt Dataframe mit einer Zeilemap2_dfrggplot() +
geom_smooth(data = rd, mapping = aes(x = MESS_DATUM, y = RS, color = STATION), size = 0.2) +
geom_abline(
data = rd_lineare_trends_2, mapping = aes(intercept = ALPHA, slope = BETA, color = STATION)
)
mk_dataframe <- function(s, df) {
t <- mk.test(na.omit(df$RS))
tibble(STATION = s, p = t$p.value)
}
rd_mk <- map2_dfr(rd_stationen$STATION, rd_stationen$data, mk_dataframe)
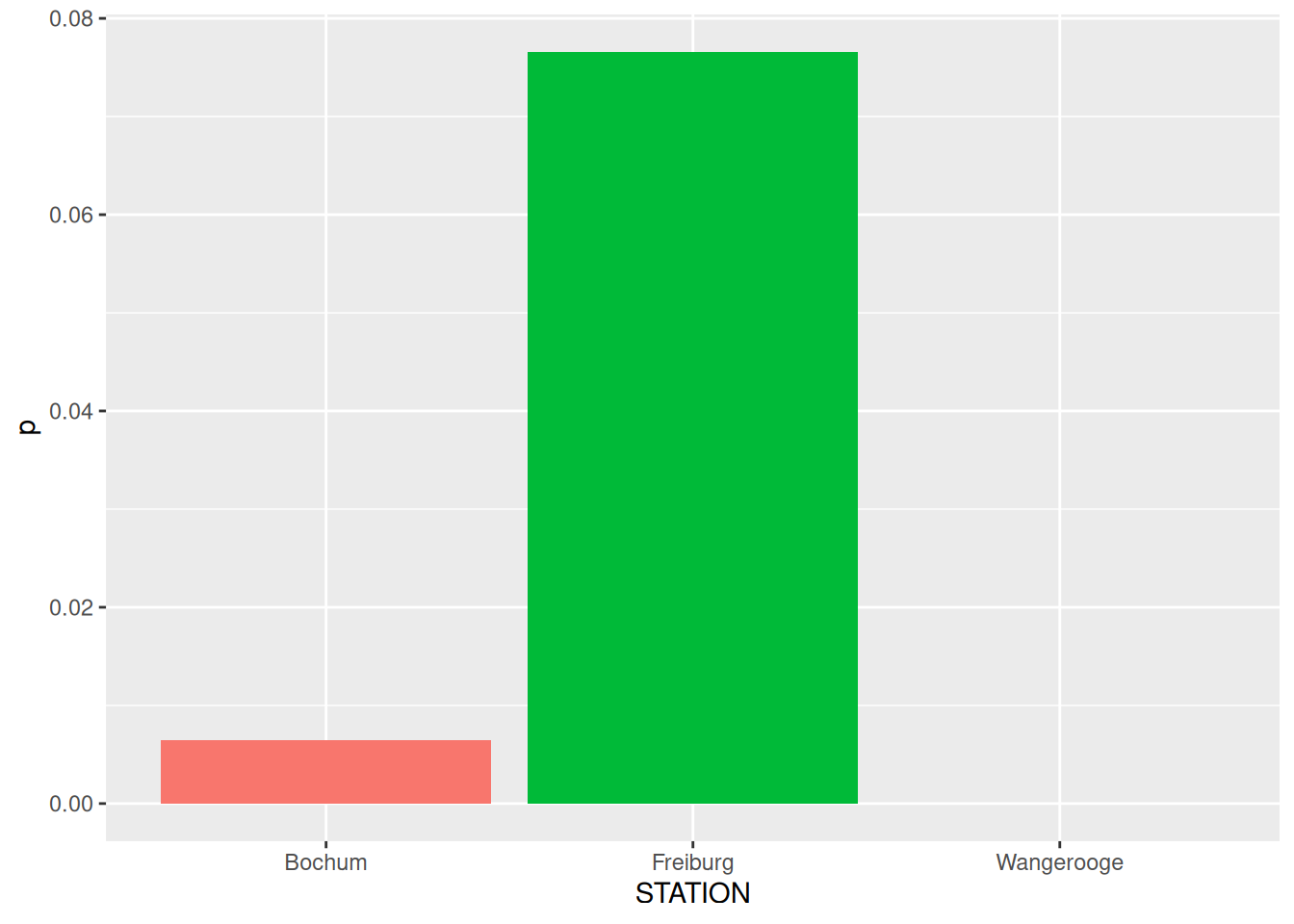
rd_mk# A tibble: 3 × 2
STATION p
<chr> <dbl>
1 Wangerooge 1.51e-10
2 Freiburg 7.66e- 2
3 Bochum 6.43e- 3
mk_test() verarbeitet keine NAsggplot(data = rd_mk) +
geom_col(mapping = aes(x = STATION, y = p, fill = STATION), show.legend = FALSE)